FM22,regression trees,steven,coomans,thesis,per2maand | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| *Unverified author* | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| R Software Module: /rwasp_regression_trees.wasp (opens new window with default values) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
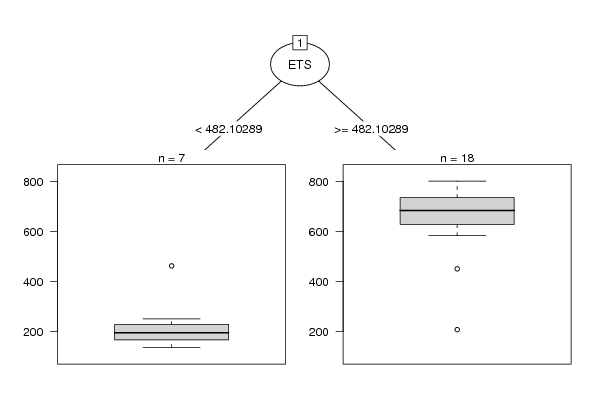
| Title produced by software: Recursive Partitioning (Regression Trees) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Date of computation: Wed, 26 May 2010 11:40:38 +0000 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cite this page as follows: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Statistical Computations at FreeStatistics.org, Office for Research Development and Education, URL http://www.freestatistics.org/blog/date/2010/May/26/t12748740771ybxyrghj8ip37w.htm/, Retrieved Wed, 26 May 2010 13:41:20 +0200 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BibTeX entries for LaTeX users: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
@Manual{KEY,
author = {{YOUR NAME}},
publisher = {Office for Research Development and Education},
title = {Statistical Computations at FreeStatistics.org, URL http://www.freestatistics.org/blog/date/2010/May/26/t12748740771ybxyrghj8ip37w.htm/},
year = {2010},
}
@Manual{R,
title = {R: A Language and Environment for Statistical Computing},
author = {{R Development Core Team}},
organization = {R Foundation for Statistical Computing},
address = {Vienna, Austria},
year = {2010},
note = {{ISBN} 3-900051-07-0},
url = {http://www.R-project.org},
}
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Original text written by user: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IsPrivate? | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| No (this computation is public) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| User-defined keywords: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FM22,regression trees,steven,coomans,thesis,per2maand | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Dataseries X: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| » Textbox « » Textfile « » CSV « | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 724 NA 726.178301126702 723.250932184245 763,275 762.275 724 724.988015993228 708.951659916233 836,3 721.125 727.8275 745.362675029711 709.122505687033 825,15 653.275 727.15725 732.118526227605 688.027750893318 764,65 663.7125 719.769025 689.036201204222 650.13316612633 820,15 735.5125 714.1633725 675.198617089505 630.378745649382 781 628.1375 716.29828525 708.15582272578 643.244311473 746,9 792.55 707.482206725 664.431555782846 612.847273246156 441,5 636.5 715.9889860525 734.439084864176 652.317786039178 1532,25 800.825 708.04008744725 680.922408372177 621.589491648231 684,75 728.05 717.318578702525 746.440564025738 660.842815301877 849,55 618.2625 718.391720832272 736.39144148354 659.888667071522 774,05 450.625 708.378798749045 671.84245826112 619.88373918091 595,525 767.525 682.60341887414 550.963003806743 534.087282375116 789 675.65 691.095576986727 669.298582567855 592.774139057074 778,45 583.25 689.551019288054 672.769176074555 597.440724777523 700,5 690.7875 678.920917359249 623.8533750 etc... | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Output produced by software: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
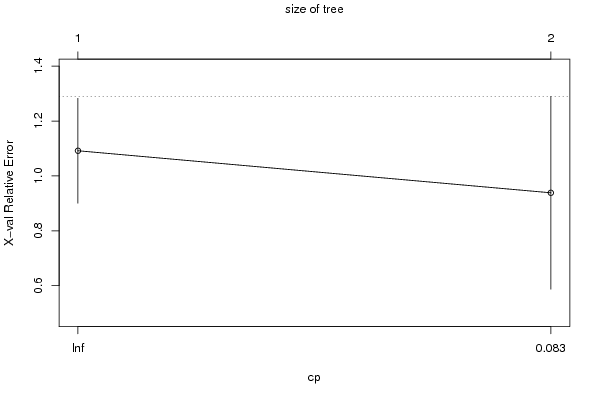
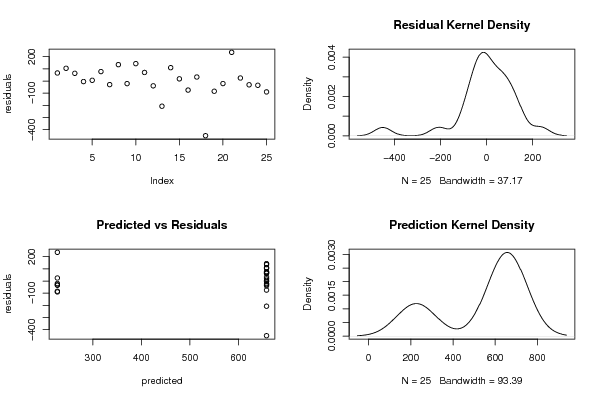
| Charts produced by software: |
| Parameters (Session): | par1 = 1 ; par2 = No ; | | Parameters (R input): | par1 = 1 ; par2 = No ; | | R code (references can be found in the software module): | library(rpart)
| library(partykit) par1 <- as.numeric(par1) autoprune <- function ( tree, method='Minimum CV'){ xerr <- tree$cptable[,'xerror'] cpmin.id <- which.min(xerr) if (method == 'Minimum CV Error plus 1 SD'){ xstd <- tree$cptable[,'xstd'] errt <- xerr[cpmin.id] + xstd[cpmin.id] cpSE1.min <- which.min( errt < xerr ) mycp <- (tree$cptable[,'CP'])[cpSE1.min] } if (method == 'Minimum CV') { mycp <- (tree$cptable[,'CP'])[cpmin.id] } return (mycp) } conf.multi.mat <- function(true, new) { if ( all( is.na(match( levels(true),levels(new) ) )) ) stop ( 'conflict of vector levels') multi.t <- list() for (mylev in levels(true) ) { true.tmp <- true new.tmp <- new left.lev <- levels (true.tmp)[- match(mylev,levels(true) ) ] levels(true.tmp) <- list ( mylev = mylev, all = left.lev ) levels(new.tmp) <- list ( mylev = mylev, all = left.lev ) curr.t <- conf.mat ( true.tmp , new.tmp ) multi.t[[mylev]] <- curr.t multi.t[[mylev]]$precision <- round( curr.t$conf[1,1] / sum( curr.t$conf[1,] ), 2 ) } return (multi.t) } x <- t(y) k <- length(x[1,]) n <- length(x[,1]) x1 <- cbind(x[,par1], x[,1:k!=par1]) mycolnames <- c(colnames(x)[par1], colnames(x)[1:k!=par1]) colnames(x1) <- mycolnames #colnames(x)[par1] m <- rpart(as.data.frame(x1)) par2 if (par2 != 'No') { mincp <- autoprune(m,method=par2) print(mincp) m <- prune(m,cp=mincp) } m$cptable bitmap(file='test1.png') plot(as.party(m),tp_args=list(id=FALSE)) dev.off() bitmap(file='test2.png') plotcp(m) dev.off() cbind(y=m$y,pred=predict(m),res=residuals(m)) myr <- residuals(m) myp <- predict(m) bitmap(file='test4.png') op <- par(mfrow=c(2,2)) plot(myr,ylab='residuals') plot(density(myr),main='Residual Kernel Density') plot(myp,myr,xlab='predicted',ylab='residuals',main='Predicted vs Residuals') plot(density(myp),main='Prediction Kernel Density') par(op) dev.off() load(file='createtable') a<-table.start() a<-table.row.start(a) a<-table.element(a,'Model Performance',6,TRUE) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'#',header=TRUE) a<-table.element(a,'Complexity',header=TRUE) a<-table.element(a,'split',header=TRUE) a<-table.element(a,'relative error',header=TRUE) a<-table.element(a,'CV error',header=TRUE) a<-table.element(a,'CV S.D.',header=TRUE) a<-table.row.end(a) for (i in 1:length(m$cptable[,1])) { a<-table.row.start(a) a<-table.element(a,i,header=TRUE) a<-table.element(a,round(m$cptable[i,'CP'],3)) a<-table.element(a,m$cptable[i,'nsplit']) a<-table.element(a,round(m$cptable[i,'rel error'],3)) a<-table.element(a,round(m$cptable[i,'xerror'],3)) a<-table.element(a,round(m$cptable[i,'xstd'],3)) a<-table.row.end(a) } a<-table.end(a) table.save(a,file='mytable.tab') | Copyright
Software written by Ed van Stee & Patrick Wessa Disclaimer Information provided on this web site is provided "AS IS" without warranty of any kind, either express or implied, including, without limitation, warranties of merchantability, fitness for a particular purpose, and noninfringement. We use reasonable efforts to include accurate and timely information and periodically update the information, and software without notice. However, we make no warranties or representations as to the accuracy or completeness of such information (or software), and we assume no liability or responsibility for errors or omissions in the content of this web site, or any software bugs in online applications. Your use of this web site is AT YOUR OWN RISK. Under no circumstances and under no legal theory shall we be liable to you or any other person for any direct, indirect, special, incidental, exemplary, or consequential damages arising from your access to, or use of, this web site. Privacy Policy We may request personal information to be submitted to our servers in order to be able to:
We NEVER allow other companies to directly offer registered users information about their products and services. Banner references and hyperlinks of third parties NEVER contain any personal data of the visitor. We do NOT sell, nor transmit by any means, personal information, nor statistical data series uploaded by you to third parties.
We store a unique ANONYMOUS USER ID in the form of a small 'Cookie' on your computer. This allows us to track your progress when using this website which is necessary to create state-dependent features. The cookie is used for NO OTHER PURPOSE. At any time you may opt to disallow cookies from this website - this will not affect other features of this website. We examine cookies that are used by third-parties (banner and online ads) very closely: abuse from third-parties automatically results in termination of the advertising contract without refund. We have very good reason to believe that the cookies that are produced by third parties (banner ads) do NOT cause any privacy or security risk. FreeStatistics.org is safe. There is no need to download any software to use the applications and services contained in this website. Hence, your system's security is not compromised by their use, and your personal data - other than data you submit in the account application form, and the user-agent information that is transmitted by your browser - is never transmitted to our servers. As a general rule, we do not log on-line behavior of individuals (other than normal logging of webserver 'hits'). However, in cases of abuse, hacking, unauthorized access, Denial of Service attacks, illegal copying, hotlinking, non-compliance with international webstandards (such as robots.txt), or any other harmful behavior, our system engineers are empowered to log, track, identify, publish, and ban misbehaving individuals - even if this leads to ban entire blocks of IP addresses, or disclosing user's identity. | ||||||||||||||||||||||||||||||||||||||||||||||