B521,regression tree,steven,coomans,thesis,per2maand | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| *Unverified author* | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| R Software Module: /rwasp_regression_trees.wasp (opens new window with default values) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
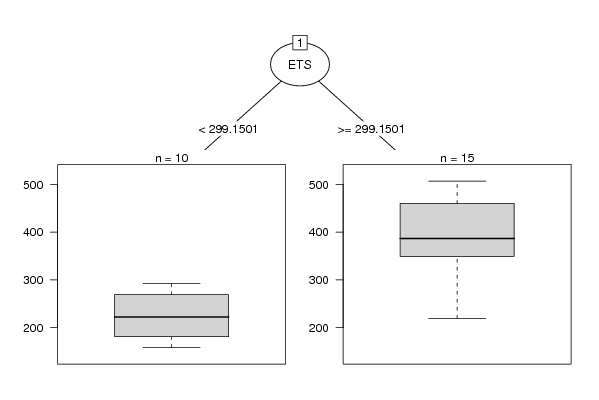
| Title produced by software: Recursive Partitioning (Regression Trees) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Date of computation: Wed, 26 May 2010 11:26:45 +0000 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cite this page as follows: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Statistical Computations at FreeStatistics.org, Office for Research Development and Education, URL http://www.freestatistics.org/blog/date/2010/May/26/t1274873276vj2ucpnjh4tu3fn.htm/, Retrieved Wed, 26 May 2010 13:27:56 +0200 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BibTeX entries for LaTeX users: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
@Manual{KEY,
author = {{YOUR NAME}},
publisher = {Office for Research Development and Education},
title = {Statistical Computations at FreeStatistics.org, URL http://www.freestatistics.org/blog/date/2010/May/26/t1274873276vj2ucpnjh4tu3fn.htm/},
year = {2010},
}
@Manual{R,
title = {R: A Language and Environment for Statistical Computing},
author = {{R Development Core Team}},
organization = {R Foundation for Statistical Computing},
address = {Vienna, Austria},
year = {2010},
note = {{ISBN} 3-900051-07-0},
url = {http://www.R-project.org},
}
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Original text written by user: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IsPrivate? | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| No (this computation is public) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| User-defined keywords: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| B521,regression tree,steven,coomans,thesis,per2maand | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Dataseries X: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| » Textbox « » Textfile « » CSV « | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 341.25 NA 333.775928269088 340.90875028286 381,75 303.6875 341.25 338.492504687478 332.86089603337 417,75 357.5 337.49375 316.528510995532 325.788548906474 359,25 295.075 339.494375 342.383916967292 327.541311770725 336 386.5755 335.0524375 312.529223276755 332.035063343418 361,8 455.6625 340.20474375 359.256754833345 355.979142780996 420,275 424.926 351.750519375 420.094418776242 422.853279739822 400,9 506.751 359.0680674375 423.143429126151 447.538297256219 248,15 433.9 373.83636069375 475.904696528962 463.452634199947 480,125 466.3375 379.842724624375 449.397276177252 469.038817658043 441,4 496.7 388.492202161938 460.087548237949 454.838864911353 483 464.45 399.312981945744 483.19214675098 483.073307566238 477,75 385.375 405.826683751169 471.364755928644 464.126997390433 486,25 381.875 403.781515376053 417.100190422978 429.475306145968 433,875 219.6375 401.590863838447 394.871034955396 366.780195970248 286,4 268.975 383.395527454603 284.288425042335 314.305881579131 231,875 292.2875 371.953 etc... | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Output produced by software: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
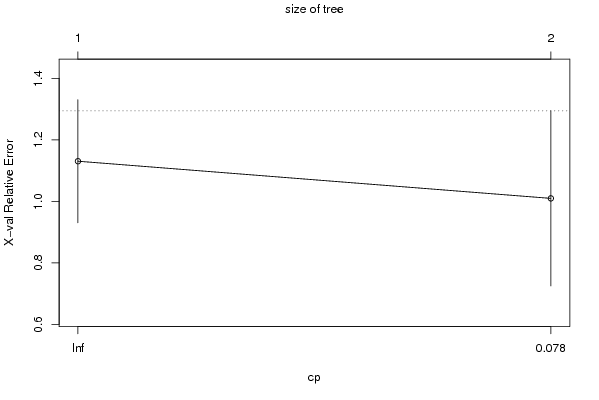
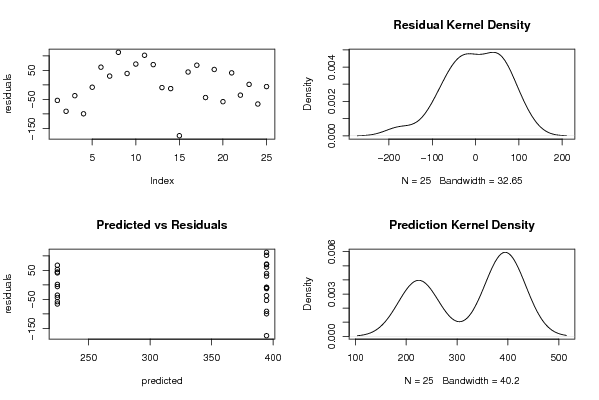
| Charts produced by software: |
| Parameters (Session): | par1 = 1 ; par2 = No ; | | Parameters (R input): | par1 = 1 ; par2 = No ; | | R code (references can be found in the software module): | library(rpart)
| library(partykit) par1 <- as.numeric(par1) autoprune <- function ( tree, method='Minimum CV'){ xerr <- tree$cptable[,'xerror'] cpmin.id <- which.min(xerr) if (method == 'Minimum CV Error plus 1 SD'){ xstd <- tree$cptable[,'xstd'] errt <- xerr[cpmin.id] + xstd[cpmin.id] cpSE1.min <- which.min( errt < xerr ) mycp <- (tree$cptable[,'CP'])[cpSE1.min] } if (method == 'Minimum CV') { mycp <- (tree$cptable[,'CP'])[cpmin.id] } return (mycp) } conf.multi.mat <- function(true, new) { if ( all( is.na(match( levels(true),levels(new) ) )) ) stop ( 'conflict of vector levels') multi.t <- list() for (mylev in levels(true) ) { true.tmp <- true new.tmp <- new left.lev <- levels (true.tmp)[- match(mylev,levels(true) ) ] levels(true.tmp) <- list ( mylev = mylev, all = left.lev ) levels(new.tmp) <- list ( mylev = mylev, all = left.lev ) curr.t <- conf.mat ( true.tmp , new.tmp ) multi.t[[mylev]] <- curr.t multi.t[[mylev]]$precision <- round( curr.t$conf[1,1] / sum( curr.t$conf[1,] ), 2 ) } return (multi.t) } x <- t(y) k <- length(x[1,]) n <- length(x[,1]) x1 <- cbind(x[,par1], x[,1:k!=par1]) mycolnames <- c(colnames(x)[par1], colnames(x)[1:k!=par1]) colnames(x1) <- mycolnames #colnames(x)[par1] m <- rpart(as.data.frame(x1)) par2 if (par2 != 'No') { mincp <- autoprune(m,method=par2) print(mincp) m <- prune(m,cp=mincp) } m$cptable bitmap(file='test1.png') plot(as.party(m),tp_args=list(id=FALSE)) dev.off() bitmap(file='test2.png') plotcp(m) dev.off() cbind(y=m$y,pred=predict(m),res=residuals(m)) myr <- residuals(m) myp <- predict(m) bitmap(file='test4.png') op <- par(mfrow=c(2,2)) plot(myr,ylab='residuals') plot(density(myr),main='Residual Kernel Density') plot(myp,myr,xlab='predicted',ylab='residuals',main='Predicted vs Residuals') plot(density(myp),main='Prediction Kernel Density') par(op) dev.off() load(file='createtable') a<-table.start() a<-table.row.start(a) a<-table.element(a,'Model Performance',6,TRUE) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'#',header=TRUE) a<-table.element(a,'Complexity',header=TRUE) a<-table.element(a,'split',header=TRUE) a<-table.element(a,'relative error',header=TRUE) a<-table.element(a,'CV error',header=TRUE) a<-table.element(a,'CV S.D.',header=TRUE) a<-table.row.end(a) for (i in 1:length(m$cptable[,1])) { a<-table.row.start(a) a<-table.element(a,i,header=TRUE) a<-table.element(a,round(m$cptable[i,'CP'],3)) a<-table.element(a,m$cptable[i,'nsplit']) a<-table.element(a,round(m$cptable[i,'rel error'],3)) a<-table.element(a,round(m$cptable[i,'xerror'],3)) a<-table.element(a,round(m$cptable[i,'xstd'],3)) a<-table.row.end(a) } a<-table.end(a) table.save(a,file='mytable.tab') | Copyright
Software written by Ed van Stee & Patrick Wessa Disclaimer Information provided on this web site is provided "AS IS" without warranty of any kind, either express or implied, including, without limitation, warranties of merchantability, fitness for a particular purpose, and noninfringement. We use reasonable efforts to include accurate and timely information and periodically update the information, and software without notice. However, we make no warranties or representations as to the accuracy or completeness of such information (or software), and we assume no liability or responsibility for errors or omissions in the content of this web site, or any software bugs in online applications. Your use of this web site is AT YOUR OWN RISK. Under no circumstances and under no legal theory shall we be liable to you or any other person for any direct, indirect, special, incidental, exemplary, or consequential damages arising from your access to, or use of, this web site. Privacy Policy We may request personal information to be submitted to our servers in order to be able to:
We NEVER allow other companies to directly offer registered users information about their products and services. Banner references and hyperlinks of third parties NEVER contain any personal data of the visitor. We do NOT sell, nor transmit by any means, personal information, nor statistical data series uploaded by you to third parties.
We store a unique ANONYMOUS USER ID in the form of a small 'Cookie' on your computer. This allows us to track your progress when using this website which is necessary to create state-dependent features. The cookie is used for NO OTHER PURPOSE. At any time you may opt to disallow cookies from this website - this will not affect other features of this website. We examine cookies that are used by third-parties (banner and online ads) very closely: abuse from third-parties automatically results in termination of the advertising contract without refund. We have very good reason to believe that the cookies that are produced by third parties (banner ads) do NOT cause any privacy or security risk. FreeStatistics.org is safe. There is no need to download any software to use the applications and services contained in this website. Hence, your system's security is not compromised by their use, and your personal data - other than data you submit in the account application form, and the user-agent information that is transmitted by your browser - is never transmitted to our servers. As a general rule, we do not log on-line behavior of individuals (other than normal logging of webserver 'hits'). However, in cases of abuse, hacking, unauthorized access, Denial of Service attacks, illegal copying, hotlinking, non-compliance with international webstandards (such as robots.txt), or any other harmful behavior, our system engineers are empowered to log, track, identify, publish, and ban misbehaving individuals - even if this leads to ban entire blocks of IP addresses, or disclosing user's identity. | ||||||||||||||||||||||||||||||||||||||||||||||