workshop 10 - recursive partitioning 5 (jonas poels) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| *Unverified author* | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| R Software Module: /rwasp_regression_trees1.wasp (opens new window with default values) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Title produced by software: Recursive Partitioning (Regression Trees) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Date of computation: Fri, 24 Dec 2010 19:26:46 +0000 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cite this page as follows: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Statistical Computations at FreeStatistics.org, Office for Research Development and Education, URL http://www.freestatistics.org/blog/date/2010/Dec/24/t1293218675628jpcrj1tq1yh4.htm/, Retrieved Fri, 24 Dec 2010 20:24:39 +0100 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BibTeX entries for LaTeX users: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
@Manual{KEY,
author = {{YOUR NAME}},
publisher = {Office for Research Development and Education},
title = {Statistical Computations at FreeStatistics.org, URL http://www.freestatistics.org/blog/date/2010/Dec/24/t1293218675628jpcrj1tq1yh4.htm/},
year = {2010},
}
@Manual{R,
title = {R: A Language and Environment for Statistical Computing},
author = {{R Development Core Team}},
organization = {R Foundation for Statistical Computing},
address = {Vienna, Austria},
year = {2010},
note = {{ISBN} 3-900051-07-0},
url = {http://www.R-project.org},
}
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Original text written by user: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IsPrivate? | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| No (this computation is public) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| User-defined keywords: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Dataseries X: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| » Textbox « » Textfile « » CSV « | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 162556 162556 1081 1081 213118 213118 230380558 6282929 1 29790 29790 309 309 81767 81767 25266003 4324047 1 87550 87550 458 458 153198 153198 70164684 4108272 0 84738 0 588 0 -26007 0 -15292116 -1212617 1 54660 54660 299 299 126942 126942 37955658 1485329 1 42634 42634 156 156 157214 157214 24525384 1779876 0 40949 0 481 0 129352 0 62218312 1367203 1 42312 42312 323 323 234817 234817 75845891 2519076 1 37704 37704 452 452 60448 60448 27322496 912684 1 16275 16275 109 109 47818 47818 5212162 1443586 0 25830 0 115 0 245546 0 28237790 1220017 0 12679 0 110 0 48020 0 5282200 984885 1 18014 18014 239 239 -1710 -1710 -408690 1457425 0 43556 0 247 0 32648 0 8064056 -572920 1 24524 24524 497 497 95350 95350 47388950 929144 0 6532 0 103 0 151352 0 15589256 1151176 0 7123 0 109 0 288170 0 31410530 790090 1 20813 20813 502 502 114337 114337 57397174 774497 1 37597 37597 248 248 37884 37884 9395232 990576 0 17821 0 373 0 122844 0 45820812 454195 1 12988 12988 119 119 82340 82340 9798460 876607 1 2 etc... | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Output produced by software: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enter (or paste) a matrix (table) containing all data (time) series. Every column represents a different variable and must be delimited by a space or Tab. Every row represents a period in time (or category) and must be delimited by hard returns. The easiest way to enter data is to copy and paste a block of spreadsheet cells. Please, do not use commas or spaces to seperate groups of digits!
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
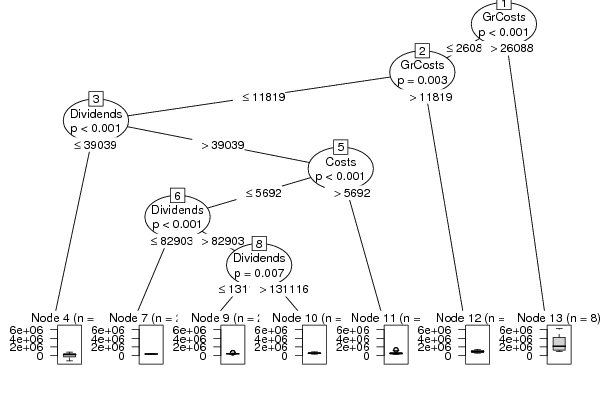
| Charts produced by software: | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Parameters (Session): | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| par1 = 9 ; par2 = none ; par3 = 3 ; par4 = no ; | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Parameters (R input): | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| par1 = 9 ; par2 = none ; par3 = 3 ; par4 = no ; | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| R code (references can be found in the software module): | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| library(party)
library(Hmisc) par1 <- as.numeric(par1) par3 <- as.numeric(par3) x <- data.frame(t(y)) is.data.frame(x) x <- x[!is.na(x[,par1]),] k <- length(x[1,]) n <- length(x[,1]) colnames(x)[par1] x[,par1] if (par2 == 'kmeans') { cl <- kmeans(x[,par1], par3) print(cl) clm <- matrix(cbind(cl$centers,1:par3),ncol=2) clm <- clm[sort.list(clm[,1]),] for (i in 1:par3) { cl$cluster[cl$cluster==clm[i,2]] <- paste('C',i,sep='') } cl$cluster <- as.factor(cl$cluster) print(cl$cluster) x[,par1] <- cl$cluster } if (par2 == 'quantiles') { x[,par1] <- cut2(x[,par1],g=par3) } if (par2 == 'hclust') { hc <- hclust(dist(x[,par1])^2, 'cen') print(hc) memb <- cutree(hc, k = par3) dum <- c(mean(x[memb==1,par1])) for (i in 2:par3) { dum <- c(dum, mean(x[memb==i,par1])) } hcm <- matrix(cbind(dum,1:par3),ncol=2) hcm <- hcm[sort.list(hcm[,1]),] for (i in 1:par3) { memb[memb==hcm[i,2]] <- paste('C',i,sep='') } memb <- as.factor(memb) print(memb) x[,par1] <- memb } if (par2=='equal') { ed <- cut(as.numeric(x[,par1]),par3,labels=paste('C',1:par3,sep='')) x[,par1] <- as.factor(ed) } table(x[,par1]) colnames(x) colnames(x)[par1] x[,par1] if (par2 == 'none') { m <- ctree(as.formula(paste(colnames(x)[par1],' ~ .',sep='')),data = x) } load(file='createtable') if (par2 != 'none') { m <- ctree(as.formula(paste('as.factor(',colnames(x)[par1],') ~ .',sep='')),data = x) if (par4=='yes') { a<-table.start() a<-table.row.start(a) a<-table.element(a,'10-Fold Cross Validation',3+2*par3,TRUE) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'',1,TRUE) a<-table.element(a,'Prediction (training)',par3+1,TRUE) a<-table.element(a,'Prediction (testing)',par3+1,TRUE) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'Actual',1,TRUE) for (jjj in 1:par3) a<-table.element(a,paste('C',jjj,sep=''),1,TRUE) a<-table.element(a,'CV',1,TRUE) for (jjj in 1:par3) a<-table.element(a,paste('C',jjj,sep=''),1,TRUE) a<-table.element(a,'CV',1,TRUE) a<-table.row.end(a) for (i in 1:10) { ind <- sample(2, nrow(x), replace=T, prob=c(0.9,0.1)) m.ct <- ctree(as.formula(paste('as.factor(',colnames(x)[par1],') ~ .',sep='')),data =x[ind==1,]) if (i==1) { m.ct.i.pred <- predict(m.ct, newdata=x[ind==1,]) m.ct.i.actu <- x[ind==1,par1] m.ct.x.pred <- predict(m.ct, newdata=x[ind==2,]) m.ct.x.actu <- x[ind==2,par1] } else { m.ct.i.pred <- c(m.ct.i.pred,predict(m.ct, newdata=x[ind==1,])) m.ct.i.actu <- c(m.ct.i.actu,x[ind==1,par1]) m.ct.x.pred <- c(m.ct.x.pred,predict(m.ct, newdata=x[ind==2,])) m.ct.x.actu <- c(m.ct.x.actu,x[ind==2,par1]) } } print(m.ct.i.tab <- table(m.ct.i.actu,m.ct.i.pred)) numer <- 0 for (i in 1:par3) { print(m.ct.i.tab[i,i] / sum(m.ct.i.tab[i,])) numer <- numer + m.ct.i.tab[i,i] } print(m.ct.i.cp <- numer / sum(m.ct.i.tab)) print(m.ct.x.tab <- table(m.ct.x.actu,m.ct.x.pred)) numer <- 0 for (i in 1:par3) { print(m.ct.x.tab[i,i] / sum(m.ct.x.tab[i,])) numer <- numer + m.ct.x.tab[i,i] } print(m.ct.x.cp <- numer / sum(m.ct.x.tab)) for (i in 1:par3) { a<-table.row.start(a) a<-table.element(a,paste('C',i,sep=''),1,TRUE) for (jjj in 1:par3) a<-table.element(a,m.ct.i.tab[i,jjj]) a<-table.element(a,round(m.ct.i.tab[i,i]/sum(m.ct.i.tab[i,]),4)) for (jjj in 1:par3) a<-table.element(a,m.ct.x.tab[i,jjj]) a<-table.element(a,round(m.ct.x.tab[i,i]/sum(m.ct.x.tab[i,]),4)) a<-table.row.end(a) } a<-table.row.start(a) a<-table.element(a,'Overall',1,TRUE) for (jjj in 1:par3) a<-table.element(a,'-') a<-table.element(a,round(m.ct.i.cp,4)) for (jjj in 1:par3) a<-table.element(a,'-') a<-table.element(a,round(m.ct.x.cp,4)) a<-table.row.end(a) a<-table.end(a) table.save(a,file='mytable3.tab') } } m bitmap(file='test1.png') plot(m) dev.off() bitmap(file='test1a.png') plot(x[,par1] ~ as.factor(where(m)),main='Response by Terminal Node',xlab='Terminal Node',ylab='Response') dev.off() if (par2 == 'none') { forec <- predict(m) result <- as.data.frame(cbind(x[,par1],forec,x[,par1]-forec)) colnames(result) <- c('Actuals','Forecasts','Residuals') print(result) } if (par2 != 'none') { print(cbind(as.factor(x[,par1]),predict(m))) myt <- table(as.factor(x[,par1]),predict(m)) print(myt) } bitmap(file='test2.png') if(par2=='none') { op <- par(mfrow=c(2,2)) plot(density(result$Actuals),main='Kernel Density Plot of Actuals') plot(density(result$Residuals),main='Kernel Density Plot of Residuals') plot(result$Forecasts,result$Actuals,main='Actuals versus Predictions',xlab='Predictions',ylab='Actuals') plot(density(result$Forecasts),main='Kernel Density Plot of Predictions') par(op) } if(par2!='none') { plot(myt,main='Confusion Matrix',xlab='Actual',ylab='Predicted') } dev.off() if (par2 == 'none') { detcoef <- cor(result$Forecasts,result$Actuals) a<-table.start() a<-table.row.start(a) a<-table.element(a,'Goodness of Fit',2,TRUE) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'Correlation',1,TRUE) a<-table.element(a,round(detcoef,4)) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'R-squared',1,TRUE) a<-table.element(a,round(detcoef*detcoef,4)) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'RMSE',1,TRUE) a<-table.element(a,round(sqrt(mean((result$Residuals)^2)),4)) a<-table.row.end(a) a<-table.end(a) table.save(a,file='mytable1.tab') a<-table.start() a<-table.row.start(a) a<-table.element(a,'Actuals, Predictions, and Residuals',4,TRUE) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'#',header=TRUE) a<-table.element(a,'Actuals',header=TRUE) a<-table.element(a,'Forecasts',header=TRUE) a<-table.element(a,'Residuals',header=TRUE) a<-table.row.end(a) for (i in 1:length(result$Actuals)) { a<-table.row.start(a) a<-table.element(a,i,header=TRUE) a<-table.element(a,result$Actuals[i]) a<-table.element(a,result$Forecasts[i]) a<-table.element(a,result$Residuals[i]) a<-table.row.end(a) } a<-table.end(a) table.save(a,file='mytable.tab') } if (par2 != 'none') { a<-table.start() a<-table.row.start(a) a<-table.element(a,'Confusion Matrix (predicted in columns / actuals in rows)',par3+1,TRUE) a<-table.row.end(a) a<-table.row.start(a) a<-table.element(a,'',1,TRUE) for (i in 1:par3) { a<-table.element(a,paste('C',i,sep=''),1,TRUE) } a<-table.row.end(a) for (i in 1:par3) { a<-table.row.start(a) a<-table.element(a,paste('C',i,sep=''),1,TRUE) for (j in 1:par3) { a<-table.element(a,myt[i,j]) } a<-table.row.end(a) } a<-table.end(a) table.save(a,file='mytable2.tab') } | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Copyright

This work is licensed under a
Creative Commons Attribution-Noncommercial-Share Alike 3.0 License.
Software written by Ed van Stee & Patrick Wessa
Disclaimer
Information provided on this web site is provided "AS IS" without warranty of any kind, either express or implied, including, without limitation, warranties of merchantability, fitness for a particular purpose, and noninfringement. We use reasonable efforts to include accurate and timely information and periodically update the information, and software without notice. However, we make no warranties or representations as to the accuracy or completeness of such information (or software), and we assume no liability or responsibility for errors or omissions in the content of this web site, or any software bugs in online applications. Your use of this web site is AT YOUR OWN RISK. Under no circumstances and under no legal theory shall we be liable to you or any other person for any direct, indirect, special, incidental, exemplary, or consequential damages arising from your access to, or use of, this web site.
Privacy Policy
We may request personal information to be submitted to our servers in order to be able to:
- personalize online software applications according to your needs
- enforce strict security rules with respect to the data that you upload (e.g. statistical data)
- manage user sessions of online applications
- alert you about important changes or upgrades in resources or applications
We NEVER allow other companies to directly offer registered users information about their products and services. Banner references and hyperlinks of third parties NEVER contain any personal data of the visitor.
We do NOT sell, nor transmit by any means, personal information, nor statistical data series uploaded by you to third parties.
We carefully protect your data from loss, misuse, alteration,
and destruction. However, at any time, and under any circumstance you
are solely responsible for managing your passwords, and keeping them
secret.
We store a unique ANONYMOUS USER ID in the form of a small 'Cookie' on your computer. This allows us to track your progress when using this website which is necessary to create state-dependent features. The cookie is used for NO OTHER PURPOSE. At any time you may opt to disallow cookies from this website - this will not affect other features of this website.
We examine cookies that are used by third-parties (banner and online ads) very closely: abuse from third-parties automatically results in termination of the advertising contract without refund. We have very good reason to believe that the cookies that are produced by third parties (banner ads) do NOT cause any privacy or security risk.
FreeStatistics.org is safe. There is no need to download any software to use the applications and services contained in this website. Hence, your system's security is not compromised by their use, and your personal data - other than data you submit in the account application form, and the user-agent information that is transmitted by your browser - is never transmitted to our servers.
As a general rule, we do not log on-line behavior of individuals (other than normal logging of webserver 'hits'). However, in cases of abuse, hacking, unauthorized access, Denial of Service attacks, illegal copying, hotlinking, non-compliance with international webstandards (such as robots.txt), or any other harmful behavior, our system engineers are empowered to log, track, identify, publish, and ban misbehaving individuals - even if this leads to ban entire blocks of IP addresses, or disclosing user's identity.