library(BEST)
par1 <- as.numeric(par1) #column number of first sample
par2 <- as.numeric(par2) #column number of second sample
par3 <- as.numeric(par3)
par4 <- as.numeric(par4)
z <- t(y)
if (par1 == par2) stop('Please, select two different column numbers')
if (par1 < 1) stop('Please, select a column number greater than zero for the first sample')
if (par2 < 1) stop('Please, select a column number greater than zero for the second sample')
if (par1 > length(z[1,])) stop('The column number for the first sample should be smaller')
if (par2 > length(z[1,])) stop('The column number for the second sample should be smaller')
if(par5=='unpaired') {
(r <- BESTmcmc(z[,par1],z[,par2], parallel=F))
}
if(par5=='paired') {
(r <- BESTmcmc(z[,par1]-z[,par2], parallel=F))
}
bitmap(file='test2.png')
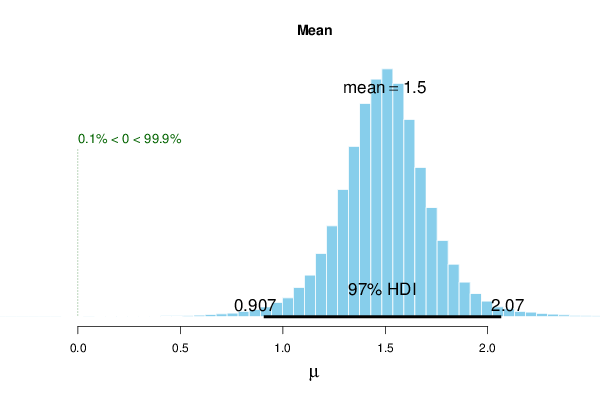
plot(r, credMass=par4, compVal=par3)
dev.off()
load(file='createtable')
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Bayesian Two Sample Test',1,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,paste('',RC.texteval('summary(r, credMass=par4, compValeff=par3)'),'',sep=''))
a<-table.row.end(a)
a<-table.end(a)
table.save(a,file='mytable.tab')
|