par1 <- as.numeric(par1) #colour
par2<- as.logical(par2) # Notches ?
par3<-as.numeric(par3) # % trim
if(par3>45){par3<-45;warning('trim limited to 45%')}
if(par3<0){par3<-0;warning('negative trim makes no sense. Trim is zero.')}
lotrm<-as.integer(length(y[1,])*par3/100)+1
hitrm<-as.integer(length(y[1,])*(100-par3)/100)
y1<-array(dim=c(dim(y)[1], hitrm-lotrm+1), dimnames=list(dimnames(y)[[1]], 1:(hitrm-lotrm+1) ))
for(i in 1:dim(y)[1]){
tmp<-order(y[i,])
y1[i,]<- y[i, tmp[lotrm:hitrm] ]
}
bitmap(file='test2.png')
pairs(t(y))
dev.off()
y<-y1
z <- as.data.frame(t(y))
bitmap(file='test1.png')
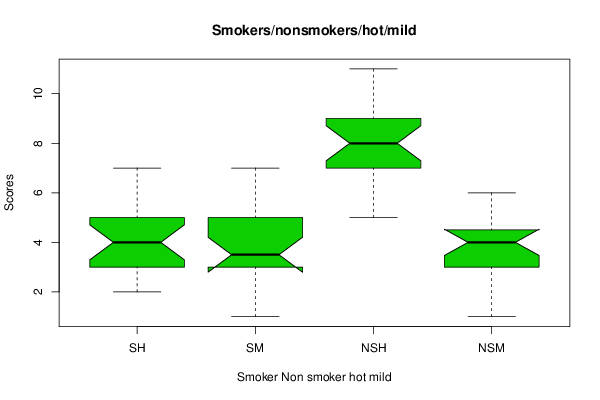
(r<-boxplot(z ,xlab=xlab,ylab=ylab,main=main,notch=par2,col=par1))
dev.off()
load(file='createtable')
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,hyperlink('overview.htm','Boxplot statistics','Boxplot overview'),6,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'Variable',1,TRUE)
a<-table.element(a,hyperlink('lower_whisker.htm','lower whisker','definition of lower whisker'),1,TRUE)
a<-table.element(a,hyperlink('lower_hinge.htm','lower hinge','definition of lower hinge'),1,TRUE)
a<-table.element(a,hyperlink('central_tendency.htm','median','definitions about measures of central tendency'),1,TRUE)
a<-table.element(a,hyperlink('upper_hinge.htm','upper hinge','definition of upper hinge'),1,TRUE)
a<-table.element(a,hyperlink('upper_whisker.htm','upper whisker','definition of upper whisker'),1,TRUE)
a<-table.row.end(a)
for (i in 1:length(y[,1]))
{
a<-table.row.start(a)
a<-table.element(a,dimnames(t(x))[[2]][i],1,TRUE)
for (j in 1:5)
{
a<-table.element(a,signif(r$stats[j,i], digits=4))
}
a<-table.row.end(a)
}
a<-table.end(a)
table.save(a,file='mytable.tab')
if (par2){
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Boxplot Notches',4,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'Variable',1,TRUE)
a<-table.element(a,'lower bound',1,TRUE)
a<-table.element(a,'median',1,TRUE)
a<-table.element(a,'upper bound',1,TRUE)
a<-table.row.end(a)
for (i in 1:length(y[,1]))
{
a<-table.row.start(a)
a<-table.element(a,dimnames(t(x))[[2]][i],1,TRUE)
a<-table.element(a,signif(r$conf[1,i], digits=4))
a<-table.element(a, signif(r$stats[3,i], digits=4))
a<-table.element(a,signif(r$conf[2,i], digits=4))
a<-table.row.end(a)
}
a<-table.end(a)
table.save(a,file='mytable1.tab')
}
a<-table.start()
a<-table.row.start(a)
a<-table.element(a,'Boxplot Means',4,TRUE)
a<-table.row.end(a)
a<-table.row.start(a)
a<-table.element(a,'Variable',1,TRUE)
a<-table.element(a,hyperlink('trimmed_mean.htm','trimmed mean','definition of trimmed mean'),1,TRUE)
a<-table.element(a,hyperlink('unbiased1.htm','unbiased SD','definition of unbiased SD'),1,TRUE)
a<-table.row.end(a)
for (i in 1:length(y[,1]))
{
a<-table.row.start(a)
a<-table.element(a,dimnames(t(x))[[2]][i],1,TRUE)
a<-table.element(a,signif(mean(z[i], trim=par3/100, na.rm=TRUE), digits=4))
a<-table.element(a,signif(sd(z[i], na.rm=TRUE), digits=4))
a<-table.row.end(a)
}
a<-table.end(a)
table.save(a,file='mytable2.tab')
|