| Multiple Linear Regression - Estimated Regression Equation |
| CorrectAnalysis[t] = + 0.0606060606060606 -0.0106060606060606T40[t] + e[t] |
| Multiple Linear Regression - Ordinary Least Squares | |||||
| Variable | Parameter | S.D. | T-STAT H0: parameter = 0 | 2-tail p-value | 1-tail p-value |
| (Intercept) | 0.0606060606060606 | 0.02914 | 2.0798 | 0.040589 | 0.020295 |
| T40 | -0.0106060606060606 | 0.060426 | -0.1755 | 0.861092 | 0.430546 |
| Multiple Linear Regression - Regression Statistics | |
| Multiple R | 0.0191475652634518 |
| R-squared | 0.000366629255518144 |
| Adjusted R-squared | -0.0115337680152494 |
| F-TEST (value) | 0.0308081526335881 |
| F-TEST (DF numerator) | 1 |
| F-TEST (DF denominator) | 84 |
| p-value | 0.861091508915969 |
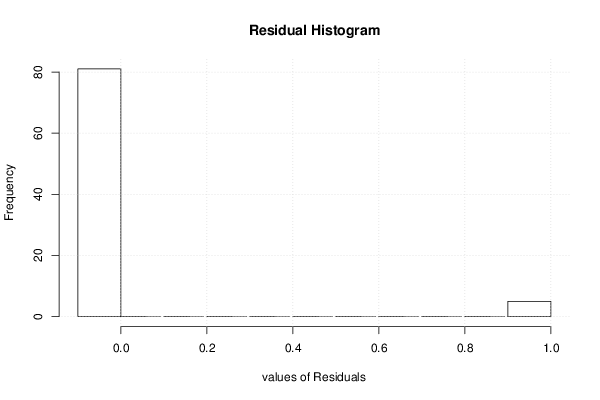
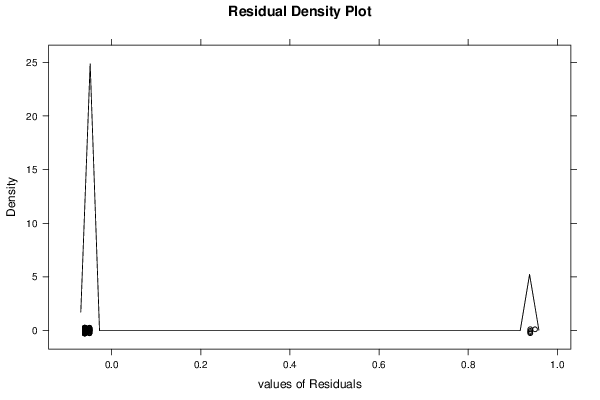
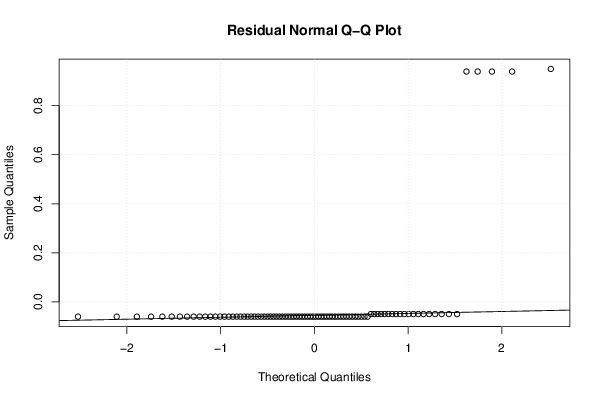
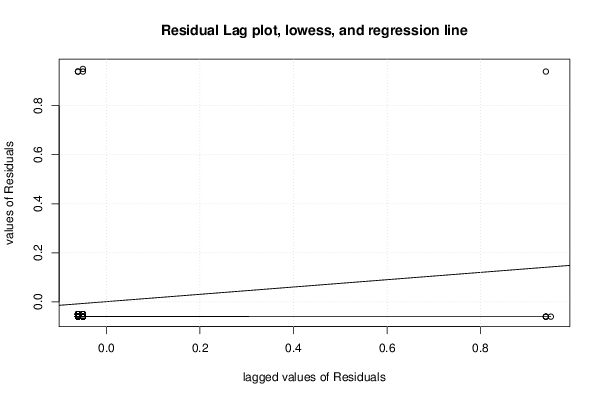
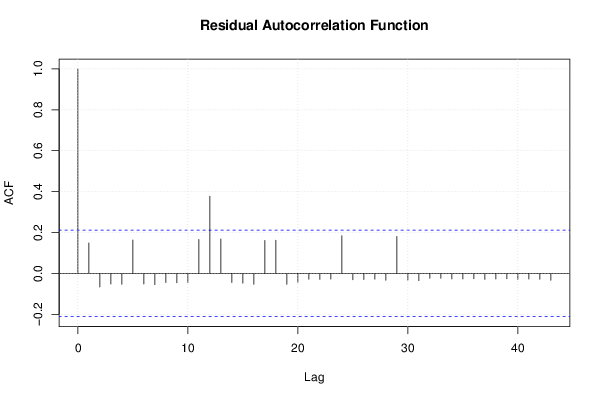
| Multiple Linear Regression - Residual Statistics | |
| Residual Standard Deviation | 0.236733116700154 |
| Sum Squared Residuals | 4.70757575757576 |
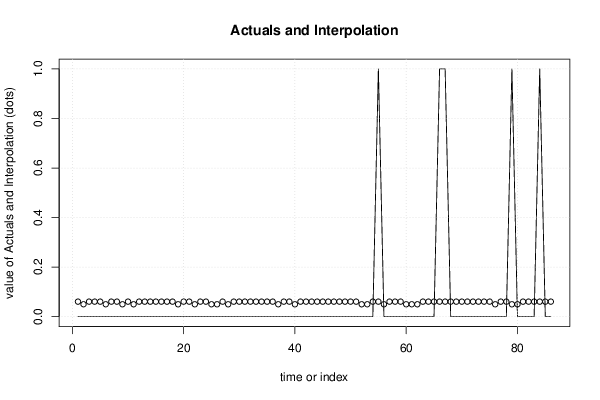
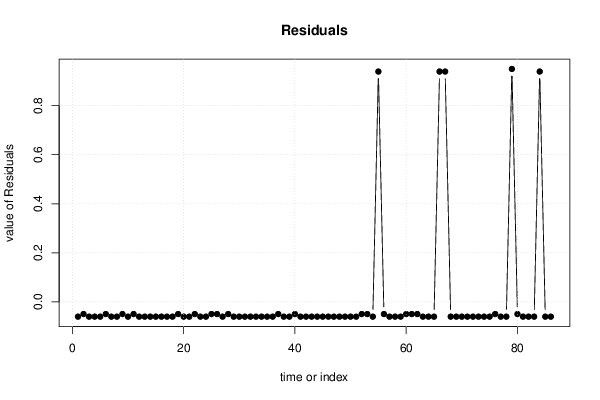
| Multiple Linear Regression - Actuals, Interpolation, and Residuals | |||
| Time or Index | Actuals | Interpolation Forecast | Residuals Prediction Error |
| 1 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 2 | 0 | 0.05 | -0.05 |
| 3 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 4 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 5 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 6 | 0 | 0.05 | -0.05 |
| 7 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 8 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 9 | 0 | 0.05 | -0.05 |
| 10 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 11 | 0 | 0.05 | -0.05 |
| 12 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 13 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 14 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 15 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 16 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 17 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 18 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 19 | 0 | 0.05 | -0.05 |
| 20 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 21 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 22 | 0 | 0.05 | -0.05 |
| 23 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 24 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 25 | 0 | 0.05 | -0.05 |
| 26 | 0 | 0.05 | -0.05 |
| 27 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 28 | 0 | 0.05 | -0.05 |
| 29 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 30 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 31 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 32 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 33 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 34 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 35 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 36 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 37 | 0 | 0.05 | -0.05 |
| 38 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 39 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 40 | 0 | 0.05 | -0.05 |
| 41 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 42 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 43 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 44 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 45 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 46 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 47 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 48 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 49 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 50 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 51 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 52 | 0 | 0.05 | -0.05 |
| 53 | 0 | 0.05 | -0.05 |
| 54 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 55 | 1 | 0.0606060606060606 | 0.939393939393939 |
| 56 | 0 | 0.05 | -0.05 |
| 57 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 58 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 59 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 60 | 0 | 0.05 | -0.05 |
| 61 | 0 | 0.05 | -0.05 |
| 62 | 0 | 0.05 | -0.05 |
| 63 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 64 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 65 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 66 | 1 | 0.0606060606060606 | 0.939393939393939 |
| 67 | 1 | 0.0606060606060606 | 0.939393939393939 |
| 68 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 69 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 70 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 71 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 72 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 73 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 74 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 75 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 76 | 0 | 0.05 | -0.05 |
| 77 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 78 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 79 | 1 | 0.05 | 0.95 |
| 80 | 0 | 0.05 | -0.05 |
| 81 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 82 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 83 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 84 | 1 | 0.0606060606060606 | 0.939393939393939 |
| 85 | 0 | 0.0606060606060606 | -0.0606060606060606 |
| 86 | 0 | 0.0606060606060606 | -0.0606060606060606 |
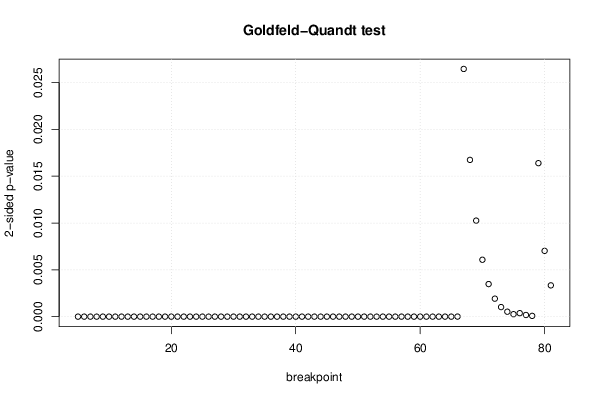
| Goldfeld-Quandt test for Heteroskedasticity | |||
| p-values | Alternative Hypothesis | ||
| breakpoint index | greater | 2-sided | less |
| 5 | 0 | 0 | 1 |
| 6 | 0 | 0 | 1 |
| 7 | 0 | 0 | 1 |
| 8 | 0 | 0 | 1 |
| 9 | 0 | 0 | 1 |
| 10 | 0 | 0 | 1 |
| 11 | 0 | 0 | 1 |
| 12 | 0 | 0 | 1 |
| 13 | 0 | 0 | 1 |
| 14 | 0 | 0 | 1 |
| 15 | 0 | 0 | 1 |
| 16 | 0 | 0 | 1 |
| 17 | 0 | 0 | 1 |
| 18 | 0 | 0 | 1 |
| 19 | 0 | 0 | 1 |
| 20 | 0 | 0 | 1 |
| 21 | 0 | 0 | 1 |
| 22 | 0 | 0 | 1 |
| 23 | 0 | 0 | 1 |
| 24 | 0 | 0 | 1 |
| 25 | 0 | 0 | 1 |
| 26 | 0 | 0 | 1 |
| 27 | 0 | 0 | 1 |
| 28 | 0 | 0 | 1 |
| 29 | 0 | 0 | 1 |
| 30 | 0 | 0 | 1 |
| 31 | 0 | 0 | 1 |
| 32 | 0 | 0 | 1 |
| 33 | 0 | 0 | 1 |
| 34 | 0 | 0 | 1 |
| 35 | 0 | 0 | 1 |
| 36 | 0 | 0 | 1 |
| 37 | 0 | 0 | 1 |
| 38 | 0 | 0 | 1 |
| 39 | 0 | 0 | 1 |
| 40 | 0 | 0 | 1 |
| 41 | 0 | 0 | 1 |
| 42 | 0 | 0 | 1 |
| 43 | 0 | 0 | 1 |
| 44 | 0 | 0 | 1 |
| 45 | 0 | 0 | 1 |
| 46 | 0 | 0 | 1 |
| 47 | 0 | 0 | 1 |
| 48 | 0 | 0 | 1 |
| 49 | 0 | 0 | 1 |
| 50 | 0 | 0 | 1 |
| 51 | 0 | 0 | 1 |
| 52 | 0 | 0 | 1 |
| 53 | 0 | 0 | 1 |
| 54 | 0 | 0 | 1 |
| 55 | 2.30546905178049e-09 | 4.61093810356098e-09 | 0.999999997694531 |
| 56 | 9.75406380728608e-10 | 1.95081276145722e-09 | 0.999999999024594 |
| 57 | 3.92431702389016e-10 | 7.84863404778032e-10 | 0.999999999607568 |
| 58 | 1.55604630640908e-10 | 3.11209261281815e-10 | 0.999999999844395 |
| 59 | 6.08904002170126e-11 | 1.21780800434025e-10 | 0.99999999993911 |
| 60 | 2.50371020076559e-11 | 5.00742040153118e-11 | 0.999999999974963 |
| 61 | 1.11954366380175e-11 | 2.23908732760349e-11 | 0.999999999988805 |
| 62 | 6.11934085445337e-12 | 1.22386817089067e-11 | 0.999999999993881 |
| 63 | 2.24479702489422e-12 | 4.48959404978845e-12 | 0.999999999997755 |
| 64 | 8.15038290163845e-13 | 1.63007658032769e-12 | 0.999999999999185 |
| 65 | 2.93707993916133e-13 | 5.87415987832267e-13 | 0.999999999999706 |
| 66 | 3.7749606503465e-06 | 7.54992130069301e-06 | 0.99999622503935 |
| 67 | 0.0132241275900069 | 0.0264482551800137 | 0.986775872409993 |
| 68 | 0.00836779927541412 | 0.0167355985508282 | 0.991632200724586 |
| 69 | 0.00512672142520157 | 0.0102534428504031 | 0.994873278574798 |
| 70 | 0.00303762151795519 | 0.00607524303591038 | 0.996962378482045 |
| 71 | 0.00173864099078952 | 0.00347728198157904 | 0.99826135900921 |
| 72 | 0.000960432467820556 | 0.00192086493564111 | 0.999039567532179 |
| 73 | 0.000511762252528802 | 0.0010235245050576 | 0.999488237747471 |
| 74 | 0.0002630871164052 | 0.0005261742328104 | 0.999736912883595 |
| 75 | 0.000130697496023803 | 0.000261394992047605 | 0.999869302503976 |
| 76 | 0.000186382444878676 | 0.000372764889757351 | 0.999813617555121 |
| 77 | 8.85717013016456e-05 | 0.000177143402603291 | 0.999911428298698 |
| 78 | 4.09861342401273e-05 | 8.19722684802546e-05 | 0.99995901386576 |
| 79 | 0.00819449040864866 | 0.0163889808172973 | 0.991805509591351 |
| 80 | 0.00351222542795434 | 0.00702445085590867 | 0.996487774572046 |
| 81 | 0.00167073809932606 | 0.00334147619865213 | 0.998329261900674 |
| Meta Analysis of Goldfeld-Quandt test for Heteroskedasticity | |||
| Description | # significant tests | % significant tests | OK/NOK |
| 1% type I error level | 73 | 0.948051948051948 | NOK |
| 5% type I error level | 77 | 1 | NOK |
| 10% type I error level | 77 | 1 | NOK |