Free Statistics
of Irreproducible Research!
Description of Statistical Computation | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Author's title | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Author | *The author of this computation has been verified* | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| R Software Module | rwasp_histogram.wasp | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
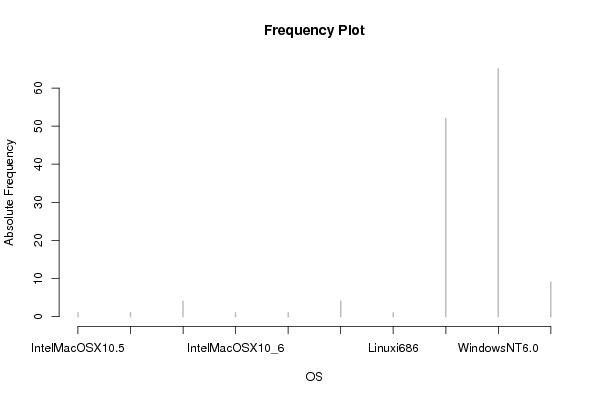
| Title produced by software | Histogram | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Date of computation | Mon, 03 Oct 2011 12:11:34 -0400 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cite this page as follows | Statistical Computations at FreeStatistics.org, Office for Research Development and Education, URL https://freestatistics.org/blog/index.php?v=date/2011/Oct/03/t1317658319hkw6hvi1e6kntb2.htm/, Retrieved Sat, 12 Jul 2025 03:30:58 +0000 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Statistical Computations at FreeStatistics.org, Office for Research Development and Education, URL https://freestatistics.org/blog/index.php?pk=125243, Retrieved Sat, 12 Jul 2025 03:30:58 +0000 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| QR Codes: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Original text written by user: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IsPrivate? | No (this computation is public) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| User-defined keywords | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Estimated Impact | 292 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Tree of Dependent Computations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Family? (F = Feedback message, R = changed R code, M = changed R Module, P = changed Parameters, D = changed Data) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| - [Histogram] [Workshop 1, task 1] [2011-10-03 16:11:34] [79818163420d1233b8d9d93d595e6c9e] [Current] - RMPD [Chi-Squared Test, McNemar Test, and Fisher Exact Test] [Taak 1 WS 6] [2011-11-11 16:49:06] [86f7284edee3dbb8ea5c7e2dec87d892] - RMPD [Chi-Squared Test, McNemar Test, and Fisher Exact Test] [Taak 1 WS 6] [2011-11-11 16:55:17] [86f7284edee3dbb8ea5c7e2dec87d892] - RMPD [Chi-Squared Test, McNemar Test, and Fisher Exact Test] [] [2011-11-11 17:11:18] [86f7284edee3dbb8ea5c7e2dec87d892] - RMPD [Chi-Squared Test, McNemar Test, and Fisher Exact Test] [] [2011-11-11 17:13:43] [86f7284edee3dbb8ea5c7e2dec87d892] | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Feedback Forum | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Post a new message | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Dataset | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Dataseries X: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'Linuxi686' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'IntelMacOSX10_6_2' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.1' 'WindowsNT6.0' 'IntelMacOSX10_6_2' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'IntelMacOSX10_5_8' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'IntelMacOSX10_5_8' 'IntelMacOSX10_5_8' 'WindowsNT6.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.0' 'IntelMacOSX10_6_2' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'IntelMacOSX10_5_7' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT5.1' 'IntelMacOSX10_6_1' 'WindowsNT6.0' 'WindowsNT5.1' 'IntelMacOSX10.5' 'WindowsNT6.0' 'IntelMacOSX10_6' 'IntelMacOSX10_5_8' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.1' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT6.0' 'WindowsNT6.1' 'WindowsNT6.1' 'WindowsNT6.1' 'WindowsNT5.1' 'WindowsNT6.0' 'WindowsNT5.1' 'IntelMacOSX10_6_2' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT5.1' 'WindowsNT5.1' | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Tables (Output of Computation) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Figures (Output of Computation) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Input Parameters & R Code | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Parameters (Session): | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| par2 = grey ; par3 = FALSE ; par4 = Unknown ; | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Parameters (R input): | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| par1 = ; par2 = grey ; par3 = FALSE ; par4 = Unknown ; | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| R code (references can be found in the software module): | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
par1 <- as.numeric(par1) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||